Through an extensive evaluation study, we report the performance of ProteinUnet2 in comparison with top SS prediction methods based on evolutionary information (SAINT and SPOT-1D). We present a lightweight deep network ProteinUnet2 for SS prediction which is based on U-Net convolutional architecture and evolutionary-based input features (from PSSM and HHblits) as well as SPOT-Contact features. Moreover, as the benchmark datasets are not random samples, the classical statistical null hypothesis testing based on the Neyman–Pearson approach is not appropriate. SS prediction as the imbalanced classification problem should not be judged by the commonly used Q3/Q8 metrics. /protein-structure-373563_final11-5c81967f46e0fb00012c667d.png)
Apart from input features calculation, the top models usually need extensive computational resources for training and prediction and are barely possible to run on a regular PC. Currently, most of the top methods use evolutionary-based input features produced by PSSM and HHblits software, although quite recently the embeddings-the new description of protein sequences generated by language models (LM) have appeared that could be leveraged as input features. As the experimental methods are expensive and sometimes impossible, many SS predictors, mainly based on different machine learning methods have been proposed for many years. 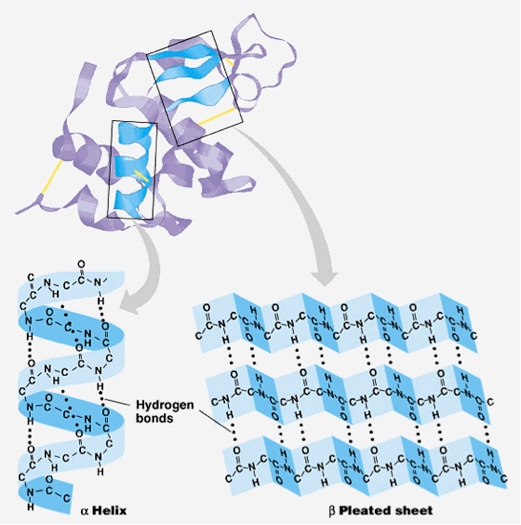
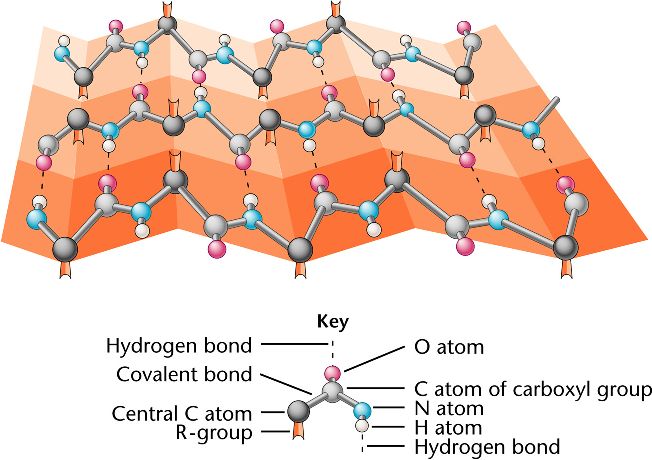
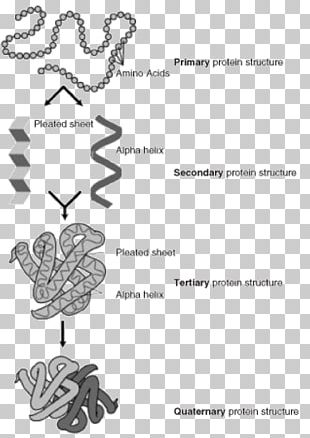
\)Ī protein’s primary structure is the unique sequence of amino acids in each polypeptide chain that makes up the protein.The prediction of protein secondary structures is a crucial and significant step for ab initio tertiary structure prediction which delivers the information about proteins activity and functions. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
